November 19, 2021
What Can Current Genetic Testing Technologies Tell You About ‘Race’?
—That It’s Still Not Real
By Nuno M. C. Martins, Michael J. Carson, and the Genetics and Society Working Group
Editor’s Note
Richard Lewontin was the co-founder of the Sociobiology Study Group of SftP Boston. The group persisted after SftP lapsed in the 1980’s, and eventually changed its name to the Genetics and Society Working Group (GSWG). GSWG has continued to monitor the periodic rise of genetic reductionism, whether it appears in the academy, industry, or popular media, to dispel its fallacy and expose the material and eugenicist consequences of biological determinism.
As Joe Graves noted, Lewontin’s elegant rebuttal of race science and prescient arguments against intentional misrepresentations of population biology has withstood the test of time, and been confirmed by new developments in population genetics. Nevertheless, those invested in racial difference are unrelenting in seizing the guise of new technologies and approaches to promote and capitalize on this social construct.
This article from members of GSWG takes us deeper into the biology of population genetics, as it seeks to address the overreach of ancestry testing. It further Graves’s article and answers the questions: How has Lewontin’s work on that issue aged? Do recent technological and scientific advances in population genetics tend to support or refute Lewontin’s conclusions? (Spoiler alert: support.)
We hope this serves as one of many primers to bring our readers up to date around the science and technology addressed in our magazine. We recommend readers to wrestle with how scientific theories have fostered racism and its political and economic aims in A History of the Concept of Race, reprinted in the July 2021 Racial Capitalism issue (Science for the People, Volume 24, No. 1).
Content At a Glance
- How do geneticists actually find genetic differences between people?
- Do people from particular continents have distinct genetic sequences from other continents, marking them as different races?
- Aren’t genome variations quite different between continents?
- When are genetic differences considered enough for organisms to be a different subspecies?
- Can genetics still allow us to separate people into distinct groups?
- Is “continental groups” just a very rough characterization?
- Can DNA ancestry tests tell me where I come from? Who are my ancestors?
- What about skin color? Isn’t that a specific genetic difference?
- So if we can’t really separate people into groups, why has our society used “race” in this way?
- What does “mixed-race” really mean?
- Is there such a thing as race on the genetic level?
Personal genetic testing is here. Comparing someone’s personal DNA sequences to those of others worldwide can reveal a shared ancestry: previously unknown similarities to other populations across the world, pointing to family history going back centuries.1
But what does that actually mean to the conversation about “race”?
The technology for decoding the human genome is now at the point where a person’s DNA sequence can be “read” for less than $1000. This has been hailed as a potential revolution in personal medicine: tailored cancer treatments, improved preventative care based on genetic predisposition to certain diseases, as well as fast diagnostics to check if a patient’s disease is affected by particular known genetic conditions.
But knowledge about genetic differences has been used before for purposes completely unrelated to healthcare. Eugenic theories have claimed superiority or inferiority of different human populations and individuals, based on genetic differences. These differences were considered proof that there are distinct “races” of people and some were intrinsically and hereditarily inferior. Although prevalent in popular discourse, these claims have no scientific basis and have been consistently debunked by biologists, anthropologists and social scientists for the past seventy years.
Yet, the lingering ideas of distinct “races”—that small differences in DNA sequence should separate people into groups and dictate a person’s human potential and their place in society—have not left us. In recent years, commercial ancestry testing has provided grounds for renewed interest in ideas of racial essentialism: that there are inherent biological or genetic traits that define all members of a racial category and are informative about who they are as people.2 Some individuals in extremist online groups are even treating their ancestry results (and the “science” behind them) as evidence of racial purity, when in reality that is not what the tests mean at all.3
In this article, we offer an overview of the science behind human genetic variation around the world. We explain how the data have been interpreted to help increase our understanding between genetics and ancestry, and how easy it can be to misinterpret the conclusions, which bolster racial essentialist arguments.
We hope that a better understanding of the nuance of genetic difference at the population level—and how migrations and geography influence these differences—will dispel the idea that race has a biological basis. We hope to make clear that race is a social construct that continues to be used to divide, discriminate, and oppress.
How do geneticists actually find genetic differences among people?
From sequencing and comparing many human genomes, it has been estimated that any two humans have an exact match of roughly 99.4% of their DNA sequence between each other.4 The remaining differences are small variations in the sequence of the DNA, and it is important to say that only a small proportion of these (9.4–11.5%) actually affect visible physical characteristics, susceptibility to disease, or other physiological traits. The rest are in fact genetically neutral: they do not have major effects on human physiology.5 Therefore, these sequences are not restricted by natural selection, and so, variations in the DNA sequence accumulate in these regions by natural random mutations, slowly over thousands of years, across individuals in a population.
Only a small portion of these DNA sequence variations are usually examined for commercial genetic testing, and they mostly sit in these genetically neutral DNA regions. The longer two human populations have been separated, the more time would mutations be allowed to accumulate and the more of these neutral differences the two populations will have. But how much these differences define separate groups has been a topic of much misunderstanding.
In 2002, a groundbreaking work by Rosenberg et al, followed by that of Bamshad et al in 2003 and Li et al in 2008, showed that using the power of DNA sequencing and statistics, they could find patterns of DNA variations across human populations from different parts of the world.6 This showed that populations native to the same continent were slightly more genetically similar to each other than to populations from other regions (figure 1A). And, at face value, this may seem to imply that individuals native to different parts of the world indeed could be considered “genetically distinct groups.” But let’s dig a little deeper into the data.
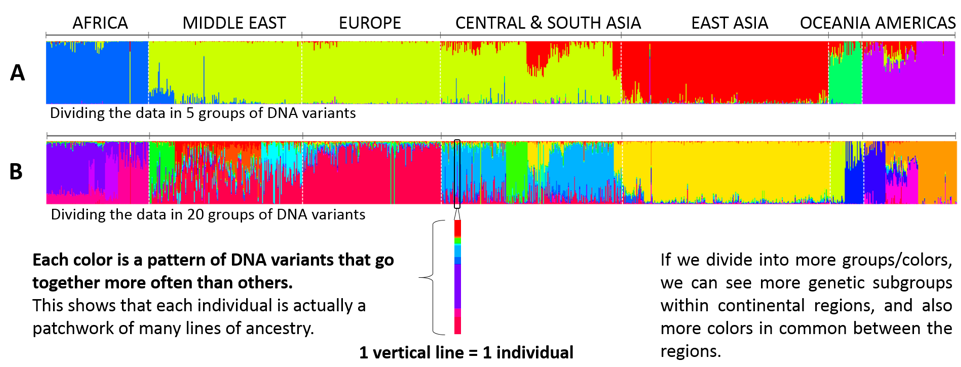
Figure 1. Substructure of the ancestry within the genome of individuals. This is the same statistical analysis that was first used by Rosenberg et al to show that populations from different continents have different patterns of DNA variants. The STRUCTURE algorithm was asked to cluster individuals into five groups of similar genetic substructure (assigning a color to each), and it turns out this coincided mostly with the individual’s actual continental origin (A). These groups of individuals have been interpreted as having more similar ancestry.7 For some individuals, most of their DNA variants belonged to a single group (ie. a single color) but for many others, part of their DNA variants also belonged to one or more of the other groups (more colors in the same individual). This tells us each individual shares partial genetic similarity with other individuals “outside” their group. We also show here that, using exactly the same genetic data used in Rosenberg’s 2002 paper, if you change the number of groups you wish to fit the individuals in, the output from STRUCTURE changes a lot: it subdivides the individuals in new groups (See figure 1B).8 The subdivisions change both within each continent (you start finding many new internal sub-groups) but also in common between continents (there is always a bit of all the other colors in every individual, at the top and bottom of each vertical line). We also know that using more or fewer individuals from each group changes the variation in the analysis and makes groups of different sizes, and causes the STRUCTURE algorithm to assign its clusters very differently, and no longer splits populations from across the world into the major continental groups.9 So these studies may still be using a number of individuals that is too low to be representative, or may even not have sufficient sub-populations of each continent to accurately represent genetic diversity across geographic distances.
Back to Top
Does that mean people from particular continents have distinct genetic sequences from other continents, marking them as different races?
No, the devil is in the details. The patterns they found are not that populations native to each continent have their own specific DNA sequence variants. Instead, the patterns are overall percentage estimates of many DNA variants in the whole population of the continent, that together form an average pattern: some variants are more common, others are more rare.
The vast majority of these DNA variants that make up the patterns are incredibly common around the world: it is just how widespread they are in each region that varies. Even those very rare DNA variants exclusive to one single population are not present in the DNA of everyone in that population.10 This means that each person within a population has a pattern of DNA variants different from the average pattern found for their population via statistical analysis (in figure 1, each vertical line is a single person).
These average patterns only actually emerge when you put together several people and look for similarities, and even then they are a statistical approximation. So if you only have one individual from each continent to work from, you could not separate them by continent: each individual is not genetically representative at all of their population’s pattern, because they would only have some of their variants matching the pattern.11 You would need several individuals from the same population to start finding some common pattern for that population (and how many individuals you choose also has an impact on the algorithm!)12 The opposite is also true: you cannot guess the full genetics of an individual just by knowing their average genetic pattern, or in other words, where their ancestors came from. You may guess at the likelihood of an individual having a certain few very specific DNA variants if you know their population(s) of origin, but even then it is just a probability.
Each individual is not genetically representative at all of their population’s genetics.
You cannot guess the genetics of an individual just by knowing where their ancestors came from.
The authors of the studies were clear in explaining their method, as well as highlighting that the differences are found in the estimated patterns of percentages of DNA variants in each population, and not specific DNA variants that distinguish populations from each other. Despite this, and because the articles are worded in technical language, it did not stop many misreadings of these works, both by experts and non-experts, as providing genetic evidence for racial essentialism to various degrees, which has wide-ranging problematic ramifications.13
Incorrect interpretation of studies like these has serious medical implications: just assuming someone from a certain part of the world is likely to have particular genetic conditions, without testing if they actually have the DNA variants associated with disease risk, may lead to incorrect diagnoses.14 For example, the genomes of African-Americans have on average 24% European ancestry, and those of European-Americans have roughly 4% of African ancestry. Assuming someone is from a certain “race,” especially when based on a few visible characteristics such as skin color, does not at all determine genetic dispositions.
Assuming someone is from a certain “race” doesn’t mean genetic conditions […] statistically associated with another “race” cannot be present in them.
The majority of the DNA variants that are associated with the different patterns are actually common to all populations. Only the percentage patterns (how common or how rare DNA variants are) change from one continental group to another (figure 2). Because so few variants are specific to a single population—and even these are not widespread across that population itself—there is not a specific set of DNA letters that genetically defines a “racial” population (that is, carried by every individual in it). What we can detect are patterns of how frequent DNA variants are across a population, compared to other populations. In this way, comparing many rarer variants in each population helps establish the likelihood of an individual’s ancestry.
There isn’t a specific set of DNA letters that genetically defines a “racial” population, carried by every individual in it.
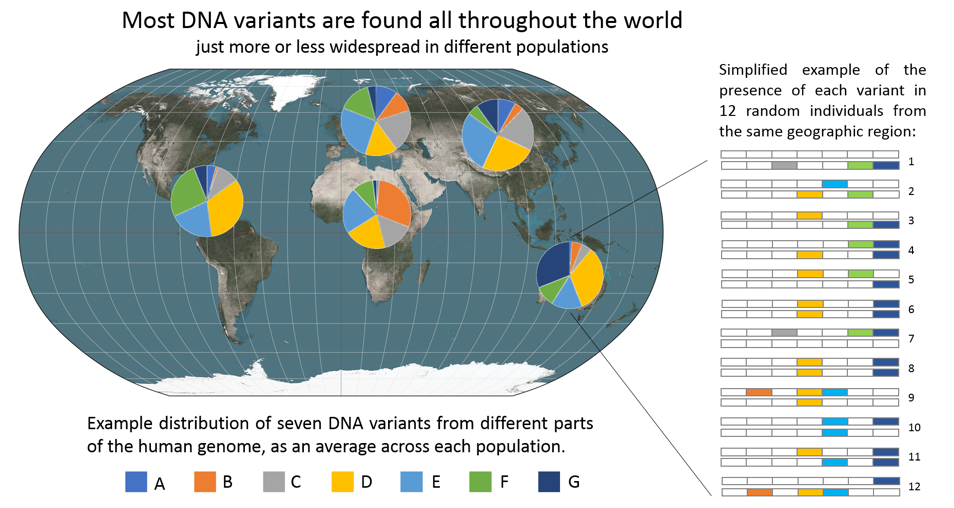
Figure 2. Most DNA variants are found all throughout the world. The figure above is an example of how differences in DNA variants (either a single DNA letter, or short combinations of letters) can be distributed across world populations. Those DNA variants that have distinct distributions in different world populations, which are used for ancestry detection, are in fact mostly shared by all world populations. But how common they are in each population varies, and many variants are rare but never fully absent in other populations. Note: each individual always has two DNA copies, one from each of their parents. Therefore each person has two chances of having any one variant.
Therefore, what the results of these studies predicted is that there is no need for genetics-based medical treatment to be different from one continental group (i.e. “race”) to another, because there are just too few genetic sequences that are specific to entire populations from a given continent.
Back to Top
But overall, the studies say each continent has a different average pattern of DNA sequences, right? Doesn’t that mean, that on average, the DNA variation in their genome is still mostly different between continents?
Again, there is a detail that is often overlooked. The majority of the DNA variation among humans is actually found within each and every population, not between them.15 Of the tiny fraction of DNA that already varies from person to person, only about 4% is actually part of the different continental patterns of DNA variants, and constitutes what can be used to calculate graphs as in figure 1. The rest, which is more than 90% of the DNA variation between all humans, is found across all individuals of the same population, and is present throughout the world in every population (figure 3).16
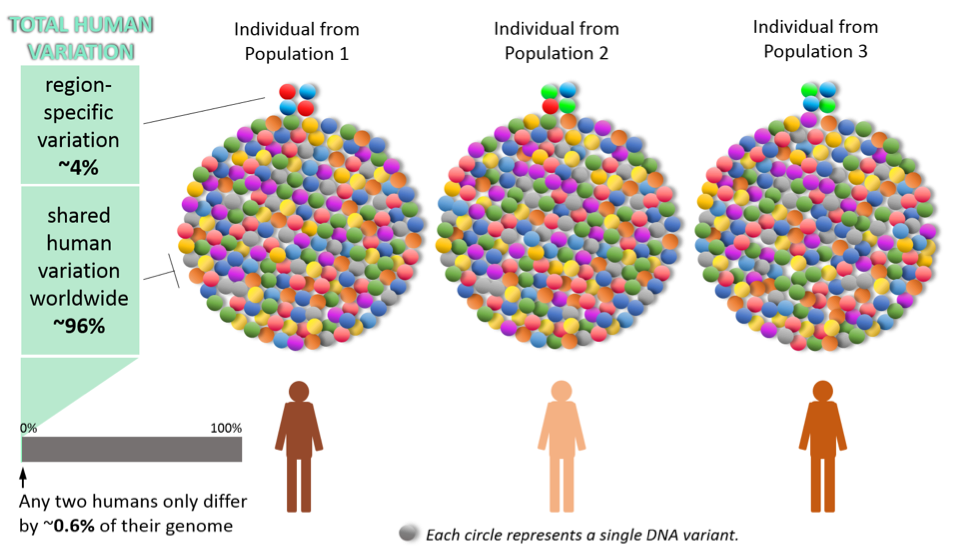
Figure 3. Scale of the DNA differences between populations of different regions of the world. Only a tiny fraction of the whole of the natural human DNA variation is actually more specific to certain geographic regions. And this total DNA variation among humans is, in itself, only a tiny fraction of the genome.
This simplified schematic in figure 3 tries to visually represent the scale of those differences. Note that the region-specific variations (the ~4% at the top) are not composed necessarily of unique DNA variants (circles of a given color) for each region, but are instead made up from different patterns of DNA variants (i.e. patterns of colors) with colors shared with other regions. In other words, different populations have several variants in common but their percentage in the population varies. What this means is that, while a person may likely share the same region-specific pattern with other people of the same population, many of their remaining DNA variants may actually be more similar to another person from across the world.
So, the variation that contributes to the DNA patterns of “different continents” (i.e. those that have been interpreted as “races”) is only a tiny fraction of the total observed variation, but the advanced statistical analysis picks up on even the smallest differences. (This ~4% of continent-specific variation is still thousands of DNA variants). The rest of the human DNA variation within each group—common among all humans—includes most physiological traits that you see manifested within every population: people have different heights, weight, big or small feet and hands, shape of their hairline, size of their noses, their cheekbones, and so on. And these are only the ones we can see: many DNA variations are physiological traits that cannot be seen outwardly (and in fact, the majority are actually neutral DNA variants). This is the common ancient variability of the human species, and all populations around the world carry it with them.
[The] mean level of difference for two individuals from the same population is almost the same as the mean level of difference for two individuals from any two populations anywhere in the world.17
It is specifically the small differences across many DNA variations, whose overall patterns vary slightly from location to location, that researchers and ancestry service companies use to estimate a person’s ancestry. It is only because they already have a big reference database as to what genetic patterns are more specific (statistically speaking) to presumed natives of each region of the world, or even specific countries, that they can compare any new individuals and see how well they match those patterns. But it is always an approximation.
This is also the reason why a customer’s ancestry results may vary from company to company; each company uses their own database for their reference populations (built in large part from their customers’ samples) and different algorithms for their analysis. And each database only consists of those DNA variants that the company considers to be the most informative; it is not analyzing all of a person’s DNA.18 Their results may also change slightly over time, as reference databases are updated. And most importantly, the statistical significance of the results decreases when trying to pinpoint ancestry similarity to the level of countries: only rough geographical regions can be approximated at reasonable confidence.
So, if a person receives the results of their commercial ancestry test saying that they are 72% English, 25% Japanese, and 3% North African, we should expect each of those values to be off by 5 to 15%. Since these components are only the current guess, that could change. If this person checks their results after a year, their previously reported North African ancestry may as well be missing, and a different set of ancestries may substitute that 3% as the database updates with more information and (hopefully) improved estimates.
Back to Top
But the genetic differences are there, right? Even if they are not all of the variation. There are different animal subspecies too: aren’t they distinct because of differences that we can see? Enough differences and you can consider it a subspecies?
There are ways to measure the overall level of difference in genetic variants between populations: this is called the Fixation index (Fst), and it has been measured in human populations. Most studied animal subspecies score an Fst of 25% or higher: chimpanzee subspecies score about 30%, for instance. But comparisons between what are considered the different human racial groups, they score no higher than 5%: humans around the world are just too closely related to each other. Today, we can say there are no living subspecies of Homo sapiens.19
Humans around the world are just too closely related to each other, our isolation is still very recent in the scale of evolution.
But most of all, let us remember that most of the differences we can even detect are from DNA regions that are neutral, which have no known effect on human physiology. So, most differences in the average DNA patterns between populations are not making us that much different from each other anyway.
Back to Top
Ok, but even if the differences are small, can genetics still allow us to separate people into distinct groups?
Not strict groups, no, and that is where the term race is completely inappropriate. Genetics may tell us how closely related populations are, and estimate that genetic difference as a number. But drawing a line separating one group from another, as if one “race” could be neatly categorized from another, is impossible.
Genetic differences accumulate slowly over geographic distances, so that very distant groups show more genetic differences (which is what these statistics pick up), but those in the middle are a mix of the two and harder to separate.20 Because people always move and mix, as they have done throughout thousands of years, boundaries between “races” cannot be found: instead you have gradients of people, not distinct groups (figure 4). People’s genetics blend into each other as you travel across the world.
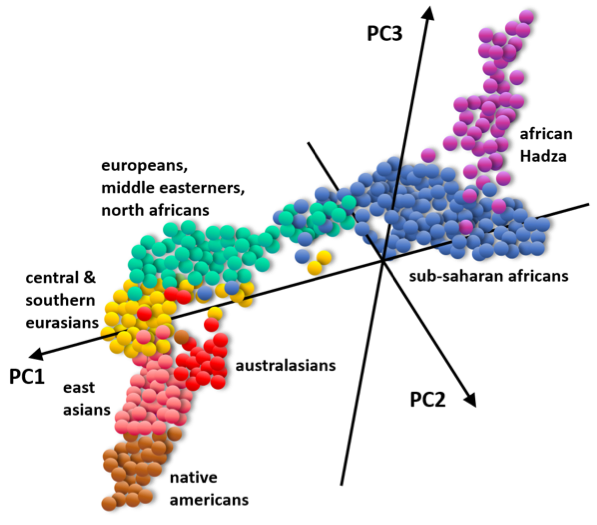
Figure 4. Populations gradually blend into each other genetically; they are never separate groups. Statistical representation of genetic similarity data from DNA samples from individuals across the world. Each dot is an individual, and they are separated or grouped depending on their degree of similarity. Each Principal Component (PC) axis is one of the three main axes of composite differences found by comparing thousands of DNA variants.21
Because many of us now live more and more in multicultural, multiethnic cities whose inhabitants can have ancestry from very distant parts of the world, more than we ever did in centuries past, it increases the impression that people look very distinct from each other if their ancestry is different. This is because many people from across the world are, in a matter of less than a century, now in the same place at the same time, in regions that have been previously more ethnically homogeneous. And thus, people visually pick up on the differences more easily. On the other hand, especially in large stretches of the world that have been continuously populated for thousands of years, such as Africa, Central Asia, South Asia, and around the Mediterranean sea, you will already find a lot of variation in people’s appearance and ethnicity as you travel across them, or even within the same city. This is particularly true in large regions where a population cannot be pinpointed to a particular “race” (as defined by any census), such as the Middle East or Central Asia, because they resemble to varying degrees all other populations in proximity. They are not “mixed-race”: this is how it has always been for thousands of years. This is why, biologically speaking, race is just a socio-political construction: a set of categories invented by humans.
The only places in the world where the genetic differences we do find become more pronounced, over shorter distances, is across difficult geographical barriers: oceans, extensive mountain ranges like the Himalayas, the Sahara, etc.22 The geographical barriers have historically isolated people from each other. (But it is important to remember that people living across the barrier were never completely isolated!) This created discontinuous jumps in genetic difference, with people becoming more similar in their genetics to one another on either side of the barrier but not across.23 This is why the different patterns between populations can be picked up by statistical analysis: it is those few patterns of DNA variants that have not homogenized across the barriers through which “groups” are created. This is especially true since most studies of worldwide genetic variation do not normally use a string of adjacent populations, but are picked from discrete points well separated from each other. Otherwise, along regions of the world that are connected, and where people have traded, travelled and mingled, genetic variation is mostly a smooth gradient, blending into the next population and the next, and the next.24
Back to Top
So, when data is presented that shows people grouped into separate “continental groups”, that is just a very rough characterization?
In a sense, yes. Plotting complex statistical data is challenging and difficult to visualize, and often researchers do so in a way that highlights what they are trying to measure, despite the reality that there is more information and a lot more nuance than can be visualized in a single graph.
So, while it is possible to analyze the degree of difference between populations on different continents and plot a graph that shows them as separated in groups, this is only a statistical representation: the “separation” is only relative. Also, many analyses often miss several “groups” that would geographically be found among the sample populations, which would “connect” the groups in a gradient of genetic variation (as shown in figure 4).
Furthermore, you cannot represent human ancestry and lineages as a tree, with groups having migrated out of Africa and spreading outwards, always separated. Human migrations have led to many groups of people mixing with other groups, travelling back to Africa, and migrations within Africa.25 An accurate description would not be a tree but the veins on a leaf: they branch and then merge again (figure 5).
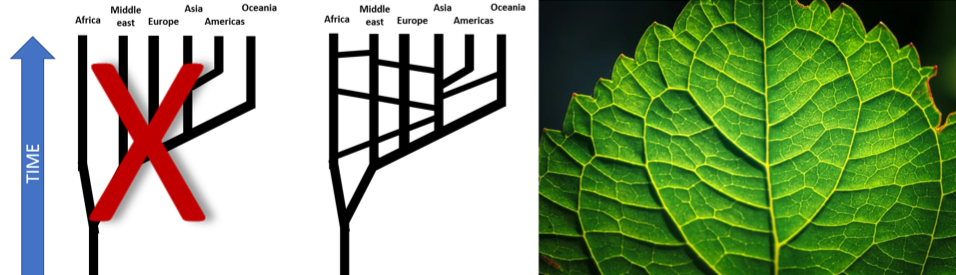
Figure 5. Human populations are not separate branches of a tree. Simplified diagram (adapted from Templeton, “Biological Races in Humans”) of the splits and joinings of populations within the human species: a tree with separate branches does not explain it, but a trellis does, which is similar to the branching and joining veins on a leaf. While different founding groups of people moved across the world and became the ancestors of the natives of each continent, the populations were never completely separate. They always mingled at the borders, and sometimes mixed together due to large migrations, already thousands of years ago before recorded history.
Back to Top
But DNA ancestry tests can still tell me where I come from, right? Who are my ancestors?
Somewhat, in an indirect way. Ancestry tests are popularly perceived to tell “where you come from,” as if they identify your ancestors within your DNA, which is not absolutely true. What it does is compare how similar you are to other people in the world of today, and then estimate the common ancestry between you and them. In a way, it is a method of genealogy, which can identify potential cousins and grandparents, but in addition, more vague and distant connections, such as shared ancestry with people from other continents.
But not all your ancestry is written in your DNA, as some parts may get lost because they are too diluted. The DNA one parent passes onto their child (which makes 50% of the child’s DNA) is a mix of the parent’s own maternal and paternal halves which come from their own parents, but which parts come from which grandparents are always random. One child may have some parts of their DNA from one grandparent, but their sibling may have those same regions of DNA coming from another grandparent. The more distant the ancestor is, the smaller these DNA parts they would receive from the ancestor, as they have been reduced by half over and over again for generations. As a result, for distant ancestries where there is only a tiny fraction of the DNA remaining from that ancestor, it may be so small that a random distribution would prevent the DNA’s passage to the next generation.
So not all of your ancestry is written in your DNA: to make better estimates of ancestry and DNA connections you always need large numbers of people to compare (because single individuals do not hold all the information).
Back to Top
What about skin color? That is something that is very strongly inherited, and varies a lot across the world. Isn’t that a specific genetic difference?
Skin color falls within those very few variations that are more specific to certain regions of the world, and that is why we see them more clearly. And there is a very good explanation why it differs so strongly, which seems contradictory when we know the majority of DNA variants between populations are mostly neutral.
Although the trait of skin color seems simple at first glance, and we use definitive words to mark differences, it is actually a complex trait based on at least 5 locations in the genome, each location with two to four actual DNA variants that can have an effect, allowing for many possible combinations. Although it seems to be a simple variation, and which has often been shown as such in diagrams, it is based on the complex interaction between multiple gene products (proteins). Think of it as an assembly line that produces a color, but how the color is blended can be tweaked at several different points (the proteins influenced by each DNA variant) along the process.
The genetics of skin color around the world actually share a common origin, and the same DNA variants that define skin color are mostly present in all populations! How can this be? While humans evolved in Africa and then spread outwards across the world, the truth is most DNA variants in people today associated with both dark and light skin have already existed in Africa for millennia. There are today still many differences in skin tones among native Africans, with some populations as light-skinned as native southern Europeans and East Asians. And dark-skinned African individuals carry DNA variants that are actually associated with light skin, but because they carry several other variants associated with dark skin, the overall color of their skin is darker. But the variants remain in the population: their effect is simply less noticeable.
Most DNA variants in people today associated with both dark and light skin already existed in Africa for millennia.
The places in the world where native individuals have darker skin are not just parts of Africa but also South Asia, Australia, Melanesia, and parts of South America: equatorial areas that experience intense ultraviolet radiation from the sun. And at latitudes adjacent to the equator, like the Mediterranean, populations show a stronger ability to tan their skin than those at higher latitudes.26 In fact, many DNA variants associated with dark skin in native Africans are also found in native South Asians and Austro-Melanesians, suggesting these DNA variants were retained in the populations as they moved out of Africa but settled in areas with equally high ultraviolet intensity.27 The very first humans to settle in northern Europe also seem to have had a pattern of DNA variants that translates to dark skin, and possibly green eyes.28
The reason why skin color seems to vary so much across the world, despite the reality that overall genetic differences are small, is believed to be environmental. Many scientists proposed that only those having the most suitable skin tone ended up surviving best in certain environments.29 As such, the difference in DNA variants, among populations in distant parts of the world, are understood to have increased faster over generations because of this adaptation pressure. Dark skin may be necessary for survival at the equator because of the high energy ultraviolet light that degrades folate through the skin, a vitamin that is essential for human fetal development.30 Light skin is believed to have become more important once humans were living at higher latitudes, for a different reason: having dark skin in those regions, where UV light is less intense, could mean not enough light to activate vitamin D production deeper in the skin. There may have been, of course, cultural and social preferences between populations that interacted with natural selection to also contribute to skin color diversification and isolation.31
So, specifically because there was strong survival selection for it, differences in skin color became one of those few traits that evolved faster than other variations in human physiology.32 It just happens to be a very visible trait to our eyes. The more intriguing question is perhaps, why, of all the visible differences among populations, do we use skin color as the most “defining” of race? Because we believe it indicates deeper genetic differences? Clearly, “genetic adaptation to sunshine” cannot support the weight of all the eugenicist claims made about differences among different human populations. More likely, it is little more than a form of shorthand that has evolved in the popular discourse to identify and refer to a specific population without much regard for actual genetic difference.
Back to Top
So if we can’t really separate people into groups, and most of the differences are tiny and superficial at the genetic level, why has our society used “race” in this way?
Because we humans like to categorize, and trying to categorize even humans has a long tradition in science.33 But all categories are arbitrary and created by us.
It is a socio-political construction, and it has been used throughout history by dominant groups of people, in a given location, as a shorthand to refer to other groups of people from places far away, either by their appearance or custom. The “race” categories change depending on the local dominant group of people or culture, and how much that group feels other groups are “different” from them.
In societies where “racial purity” has been an entrenched belief and practice, the genetics of people who live in such societies can reflect racial oppression. In the Americas, both the United States in the north and Latin American countries in the south had been parts of the Atlantic slave trade. However, the ancestry contribution in individuals of African descent today in these two regions is quite different. For individuals who have at least 5% of African ancestry (as measured by DNA variant testing as we mentioned above), a clear distinction can be seen: those from the United States have a higher degree of African ancestry than those from Latin America, despite the fact that Latin America received about 70% of all slaves between the 1500s and 1800s.34 This is consistent with the United States’s historical practice of racial segregation, in contrast to Latin America’s promotion of mixing (whose motivation was the reprehensible “whitening” of the population). In this manner, the word “race” in terms of thinking in racial categories (as opposed to simply parts of an individual’s ancestry) is a social concept that is more alive in countries where mixing has been uncommon. Furthermore, as racial integration progresses in countries with a history of segregation, much of what are considered “racial categories” in a national census start to lose their meaning for an increasing number of people.35 For them, these categories do not apply and perhaps never should have.
Back to Top
Then what does “mixed-race” really mean? Would that become a new “race” category in the census?
Maybe there should be no categories at all. What is considered “mixed-race” today has already happened many times in history, and is often overlooked when talking about “race” from its euro-centric perspective, and the racial categories popularized since the 1500s.36
The entire region of the Mediterranean Sea (exemplified by the Roman Empire, from Iberia to North Africa and the Middle East), as well as the Silk Road (cutting from the Middle East through Central Asia to China), are good examples of places with a history of exchange of goods, ideas, and people. And its peoples show that history in their genetics. But nobody talks of them as “mixed race.”
Population history affects a country’s pattern of DNA variations. Migrations (voluntary or forced) and many changes in national borders, even over the past 2000 years, essentially mean that what you may think is a country’s “natural” genetic pattern is often the result of a population mix throughout history. This is very much the case for Central and South America (ancestry from native South Americans, Iberians and West Africans), Spain and Portugal (ancestry from Southern and Northern Europe as well as the Middle East and North Africa), and the length of Afghanistan, Kazakhstan, Kyrgyzstan, Tajikistan, Uzbekistan, Turkmenistan, and Mongolia (ancestry from Central Asia, the Middle East, Northern India, and China).37 And importantly to the Eurocentric view of “races,” Europeans themselves are of mixed ancestry since the Bronze Age: between the first agricultural settlers originated from the Middle East and pastoralist migrants from Central Asia, who spread the Indo-European languages.38
If you look far enough in time, there has always been population mixing. This is something that commercial ancestry tests, while they can tell you your ancestry in common to people of other countries, are not clear in explaining whether the ancestries they list for you are themselves ancient mixes already.
There is no such thing as “race” at a genetic level: Each of us is made up of multiple mixed ancestries, some from parts of the world that are closer to you , others from more distant places, and some mixed ancestries are so ancient that nobody remembers culturally they were ever a mixture. This is all because humans move and mix together, as they always have, over time and across the millennia.
Therefore, we all have ancestries, not a single defining ancestry. Let alone a “race” category that any person can neatly be fitted in. For each of us, it is a long, milenia-spanning family tree, full of probabilities and estimates: the farther back you trace, the more blurry the path.
And even if statistics can trace our ancestral connections to one another, it will never show us that all individuals of a certain presumed “race” carry a set of specific genetic markers, whether it be for disease or any other quality.
Back to Top
—
About the Authors
Nuno M. C. Martins is a postdoctoral research fellow at Harvard Medical School, where he studies the structure of genes and chromosomes at the nanometer scale. He is a member of GSWG with a particular interest in simplifying the teaching and public awareness of difficult and misunderstood concepts related to genetics and society.
Michael J. Carson is Professor of Biological Sciences at Bridgewater State University in Massachusetts where he teaches Genetics, Human Genetics and Genomics for undergraduate students. He is a member of GSWG with interest in social issues related to human genetic variation, such as addressing misconceptions that human race is biological.
The Genetics and Society Working Group (GSWG) has worked for 30 years on anticipating the social impact of developments in human genetics and raising awareness among scientists about the impact of their work, encouraging them to interact with science educators and the media. It is composed of scientists, students, and professionals trained in a variety of disciplines, including genetics, sociology, ethics, and law. (https://genesandsociety.sites.northeastern.edu/members/).
—
References
- Deborah A. Bolnick et al., “Genetics. The Science and Business of Genetic Ancestry Testing,” Science 318, no. 5849 (October 19, 2007): 399–400, http://doi.org/10.1126/science.1150098.
- Aaron Panofsky and Joan Donovan, “Genetic Ancestry Testing Among White Nationalists: From Identity Repair to Citizen Science,” Social Studies of Science 49, no. 5 (October 2019): 653–681, https://doi.org/10.1177/0306312719861434; Wendy D. Roth et al., “Do Genetic Ancestry Tests Increase Racial Essentialism? Findings from a Randomized Controlled Trial,” PLoS ONE 15, no. 1 (January 2020): e0227399, https://doi.org/10.1371/journal.pone.0227399.
- Panofsky and Donovan, “Genetic Ancestry,” 653.
- Total DNA variation between any two individuals was estimated by the difference a typical sequenced human genome shows from the reference human genome. Total variation, on average in number of base pairs, is between five million (0.16% of the genome) if we count mostly single-nucleotide variants, and twenty-five million (0.8% of the genome) if we include larger structural variants, like duplications or deletions which mostly do not introduce new DNA variants into the mixture but change the number of copies of genes. “We find that a typical genome differs from the reference human genome at 4.1 million to 5.0 million sites […]. Although >99.9% of variants consist of SNPs and short indels, structural variants affect more bases: the typical genome contains an estimated 2,100 to 2,500 structural variants (∼1,000 large deletions, ∼160 copy-number variants, ∼915 Alu insertions, ∼128 L1 insertions, ∼51 SVA insertions, ∼4 NUMTs, and ∼10 inversions), affecting ∼20 million bases of sequence.” See 1000 Genomes Project Consortium et al., “A Global Reference for Human Genetic Variation,” Nature 526, no. 7571 (October 2015): 68–74, https://doi.org/10.1038/nature15393.
- A. M. Bowcock et al., “High Resolution of Human Evolutionary Trees with Polymorphic Microsatellites” Nature 368, no. 6470 (March 1994): 455–457, https://doi.org/10.1038/368455a0; Naoko Takezaki and Masatoshi Nei, “Genomic Drift and Evolution of Microsatellite DNAs in Human Populations,” Molecular Biology and Evolution 26, no. 8 (August, 2009): 1835–40, https://doi.org/10.1093/molbev/msp091.
- Noah A. Rosenberg et al., “Genetic Structure of Human Populations,” Science 298, no. 5602 (December 2002): 2381–5, https://doi.org/10.1126/science.1078311; Michael J Bamshad et al., “Human Population Genetic Structure and Inference of Group Membership,” American Journal of Human Genetics 72, no. 3 (March 2003): 578–89, https://doi.org/10.1086/368061; Jun Z. Li et al., “Worldwide Human Relationships Inferred from Genome-Wide Patterns of Variation,” Science 319 no. 5866 (February 2008): 1100–4, https://doi.org/10.1126/science.1153717.
- Liliana Porras-Hurtado et al., “An Overview of STRUCTURE: Applications, Parameter Settings, and Supporting Software,” Frontiers in Genetics 4 (May 29, 2013): 98, http://doi.org/10.3389/fgene.2013.00098.
- Deborah A. Bolnick, “Individual Ancestry Inference and the Reification of Race as a Biological Phenomenon,” in Revisiting Race in a Genomic Age, ed. Barbara Koenig and Sandra Soo-Jin Lee (New Brunswick, NJ: Rutgers University Press, 2008), 70–85.
- Nicholas G. Crawford et al., “Loci Associated with Skin Pigmentation Identified in African Populations” Science 358, no. 6365 (November 2017): eaan8433, https://doi.org/10.1126/science.aan8433; Erratum: Science 367, no. 6475 (January 2020): eaba7178, https://10.1126/science.aba7178.
- Rosenberg, “Genetic Structure of Human Populations,” 2381; Anders Bergström et al., “Insights into Human Genetic Variation and Population History from 929 Diverse Genomes,” Science. 367, no. 6484 (March 2020): eaay5012, https://doi.org/10.1126/science.aay5012.
- David J. Witherspoon, “Genetic Similarities Within and Between Human Populations,” Genetics 176, no. 1 (May 2007): 351–9, https://doi.org/10.1534/genetics.106.067355.
- Michael J. Bamshad et al., “Human Population Genetic Structure and Inference of Group Membership,” American Journal of Human Genetics 72, no. 3 (March 2003): 578–89, https://doi.org/10.1086/368061.
- Nicholas Wade, A Troublesome Inheritance: Genes, Race and Human History (Penguin, 2014); Charles Murray, Human Diversity: The Biology of Gender, Race, and Class (Grand Central Publishing, 2020).
- Iris Schrijver et al., “The Spectrum of CFTR Variants in Nonwhite Cystic Fibrosis Patients: Implications for Molecular Diagnostic Testing,” The Journal of Molecular Diagnostics 18, no. 1 (January 2016): 39–50, https://doi.org/10.1016/j.jmoldx.2015.07.005; Arjun K Manrai et al., “Genetic Misdiagnoses and the Potential for Health Disparities,” New England Journal of Medicine, 375, no. 7 (August 2016): 655–65, https://doi.org/10.1056/NEJMsa1507092; Sharon Begley, “Should Biologists Stop Grouping Us by Race?” StatNews, February 16, 2016, https://www.statnews.com/2016/02/04/should-geneticists-move-beyond-race.
- Richard Lewontin, “The Apportionment of Human Diversity,” in Evolutionary Biology, ed. T. Dobzhansky, M. K. Hecht, W. C. Steere (New York, NY: Springer, 1972), 381–398, https://doi.org/10.1007/978-1-4684-9063-3_14; Guido Barbujani et al., “An Apportionment of Human DNA Diversity,” Proceedings of the National Academy of Sciences 94, no. 9 (April 1997): 4516–9, https://doi.org/10.1073/pnas.94.9.4516; Rosenberg, “Genetic Structure of Human Populations,” 2381; Noah A. Rosenberg et al., “Clines, Clusters, and the Effect of Study Design on the Inference of Human Population Structure,” PLoS Genetics 1, no. 6 (December 2005): e70, https://doi.org/10.1371/journal.pgen.0010070.
- Noah A. Rosenberg et al., “Genetic Structure of Human Populations,” Science 298, no. 5602 (December 20, 2002): 2381–85, https://doi.org/10.1126/science.1078311.
- Noah A. Rosenberg, “A Population-Genetic Perspective on the Similarities and Differences Among Worldwide Human Populations,” Human Biology 83, no. 6 (December 2011): 659–84, https://doi.org/10.3378/027.083.0601.
- Sheldon Krimsky and David Cay Johnston, Ancestry DNA Testing and Privacy: A Consumer Guide, published by the Council for Responsible Genetics, https://sites.tufts.edu/sheldonkrimsky/files/2018/05/pub2017AncestryDNAPrivacy.pdf.
- Katarzyna Bryc et al., “Genome-Wide Patterns of Population Structure and Admixture in West Africans and African Americans,” Proceedings of the National Academy of Sciences 107, no. 2 (Jan 2010): 786–791, https://doi.org/10.1073/pnas.0909559107; Alan R. Templeton, “Biological Races in Humans,” Studies in History and Philosophy of Biological and Biomedical Sciences 44, no. 3 (September 2013): 262–71, https://doi.org/10.1016/j.shpsc.2013.04.010.
- Rosenberg, “Genetic Structure of Human Populations,” 2381; Rosenberg, “Clines, Clusters, and the Effect of Study Design”, e70; Templeton, “Biological Races in Humans,” 262.
- Sarah A. Tishkoff et al., “The Genetic Structure and History of Africans and African Americans,” Science 324, no. 5930 (May 2009): 1035–44, https://doi.org/10.1126/science.1172257.
- Rosenberg, “Clines, Clusters, and the Effect of Study Design”, e70; Rosenberg, “A Population-Genetic Perspective”, 659.
- Rosenberg, “Clines, Clusters, and the Effect of Study Design”, e70; Benjamin M Peter, Desislava Petkova, John Novembre, “Genetic Landscapes Reveal How Human Genetic Diversity Aligns with Geography,” Molecular Biology and Evolution 37, no. 4 (April 2020): 943–951, https://doi.org/10.1093/molbev/msz280.
- David Serre and Svante Pääbo, “Evidence for Gradients of Human Genetic Diversity Within and Among Continents,” Genome Research 14, no. 9 (September 2004): 1679–85, https://doi.org/10.1101/gr.2529604; Rosenberg, “A Population-Genetic Perspective”, 659; Peter, Petkova, Novembre, “Genetic Landscapes”, 943.
- Templeton, “Biological Races in Humans,” 262.
- Jennifer K. Wagner et al., “Skin Responses to Ultraviolet Radiation: Effects of Constitutive Pigmentation, Sex, and Ancestry,” Pigment Cell Research 15, no. 5 (October 2002): 385–90, https://doi.org/10.1034/j.1600-0749.2002.02046.x; Hongmei Nan et al., “Genome-wide Association Study Of Tanning Phenotype In A Population Of European Ancestry,” The Journal of Investigative Dermatology 129, no. 9 (September 2009): 2250–7, https://doi.org/10.1038/jid.2009.62; Ellen E Quillen, “The Evolution of Tanning Needs Its Day in the Sun,” Human Biology 87, no. 4 (October 2015): 352–360, https://doi.org/10.13110/humanbiology.87.4.0352.
- Crawford, “Loci Associated with Skin Pigmentation.”
- Selina Brace et al., “Ancient Genomes Indicate Population Replacement in Early Neolithic Britain,” Nature Ecology and Evolution 3, no. 5 (May 2019): 765–771.
- Crawford, “Loci Associated with Skin Pigmentation.”
- Nina G. Jablonski and George Chaplin, “The Evolution Of Human Skin Coloration,” Journal of Human Evolution 39, no. 1 (July 2000): 57–106, https://doi.org/10.1006/jhev.2000.0403; Patrice Jones et al., “The Vitamin D-Folate Hypothesis as an Evolutionary Model for Skin Pigmentation: An Update and Integration of Current Ideas,” Nutrients 10, no. 5 (April 2018): 554, https://doi.org/10.3390/nu10050554.
- Nina G. Jablonski, “The Evolution of Human Skin Pigmentation Involved the Interactions of Genetic, Environmental, and Cultural Variables.” Pigment Cell Melanoma Research 34 (2021): 707–729, https://doi.org/10.1111/pcmr.12976.
- Nina G. Jablonski and George Chaplin, “Colloquium Paper: Human Skin Pigmentation as an Adaptation to UV Radiation,” Proceedings of the National Academy of Sciences of the United States of America 107 Suppl 2 (May 11, 2010): 8962–68.
- The Linnean Society of London, “Linnaeus and Race,” September 3, 2020, https://www.linnean.org/learning/who-was-linnaeus/linnaeus-and-race.
- Steven J Micheletti et al., “Genetic Consequences of the Transatlantic Slave Trade in the Americas,” American Journal of Human Genetics 107, no. 2 (August 2020): 265–277, https://doi.org/10.1016/j.ajhg.2020.06.012.
- “Multiracial in America,” Pew Research Center, accessed September 20, 2021, https://www.pewsocialtrends.org/2015/06/11/multiracial-in-america/.
- Garrett Hellenthal et al., “A Genetic Atlas Of Human Admixture History,” Science, 343, no. 6172 (February 2014): 747–751, https://doi.org/10.1126/science.1243518; Rasmus Nielsen et al., “Tracing the Peopling of the World through Genomics,” Nature 541, no. 7637 (January 2017): 302–310, https://doi.org/10.1038/nature21347.
- Nielsen, “Tracing the Peopling,” 302.
- M.E. Allentoft et al., “Population Genomics of Bronze Age Eurasia,” Nature 522, no. 7555 (June 2015): 167–72, https://doi.org/10.1038/nature14507; Wolfgang Haak, et al., “Massive Migration from the Steppe was a Source for Indo-European Languages in Europe,” Nature 522, no. 7555 (June 2015): 207–11, https://doi.org/10.1038/nature14317.